![If Score(i, j) denotes best score to aligning A[1 : i] and B[1 : j] Score(i-1, j) + galign A[i] with GAP Score(i, j-1) + galign B[j] with GAP Score(i, - If Score(i, j) denotes best score to aligning A[1 : i] and B[1 : j] Score(i-1, j) + galign A[i] with GAP Score(i, j-1) + galign B[j] with GAP Score(i, -](https://images.slideplayer.com/15/4857608/slides/slide_18.jpg)
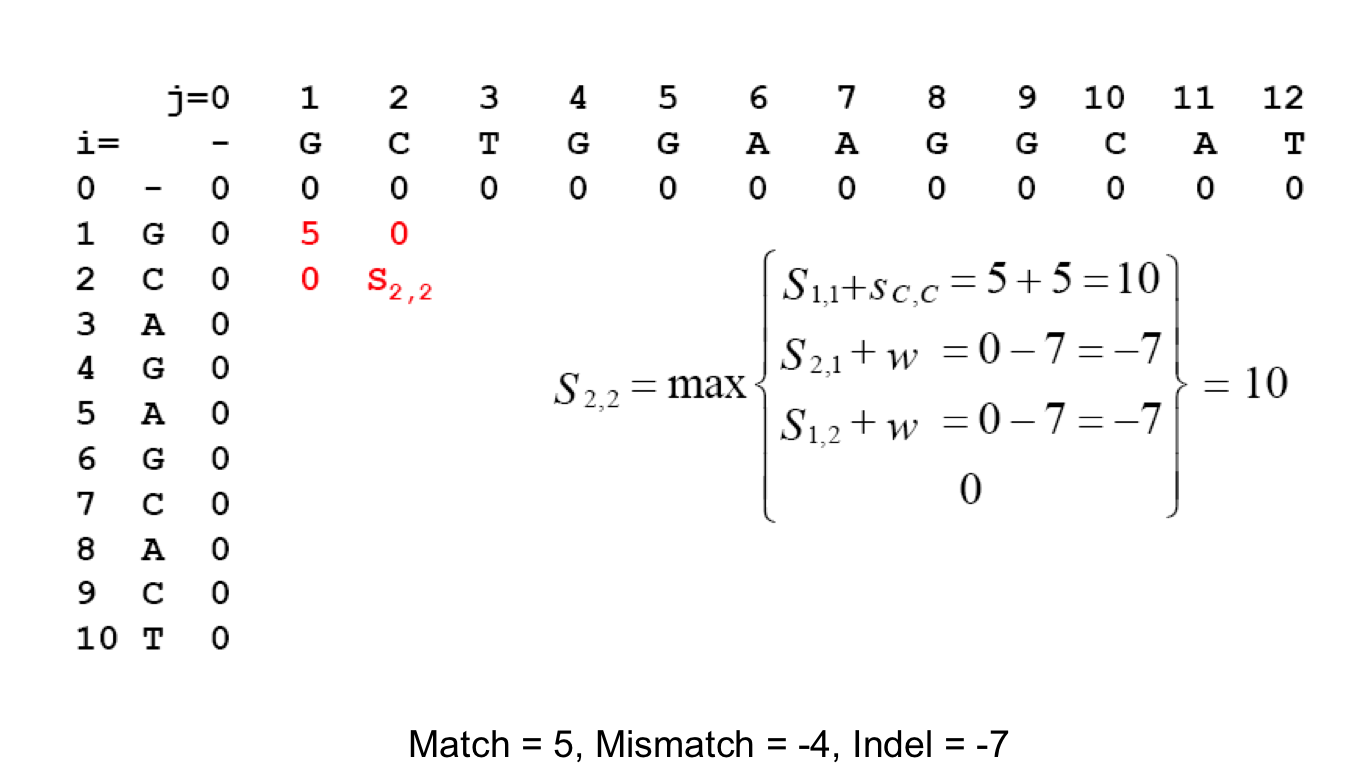
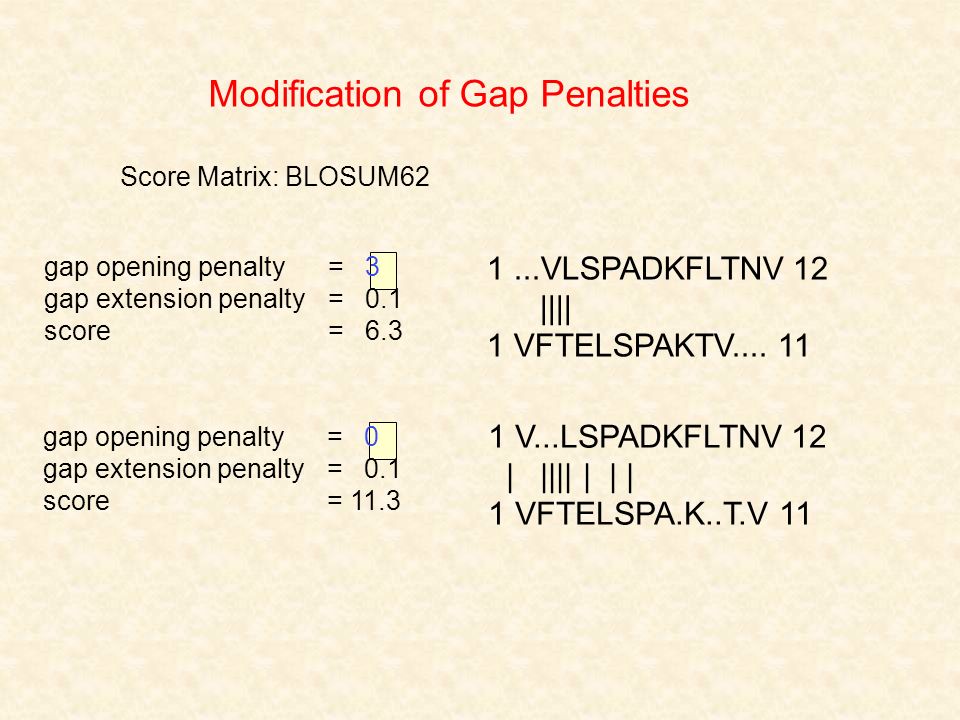
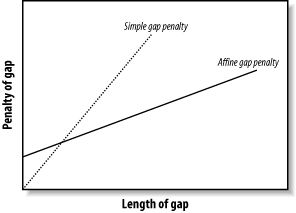
If Score(i, j) denotes best score to aligning A[1 : i] and B[1 : j] Score(i-1, j) + galign A[i] with GAP Score(i, j-1) + galign B[j] with GAP Score(i, -
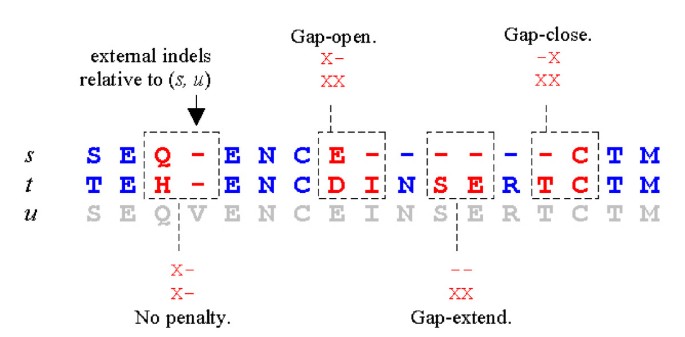
MUSCLE: a multiple sequence alignment method with reduced time and space complexity | BMC Bioinformatics | Full Text

Position-specific gap penalties. An alignment of two profiles X and Y.... | Download Scientific Diagram
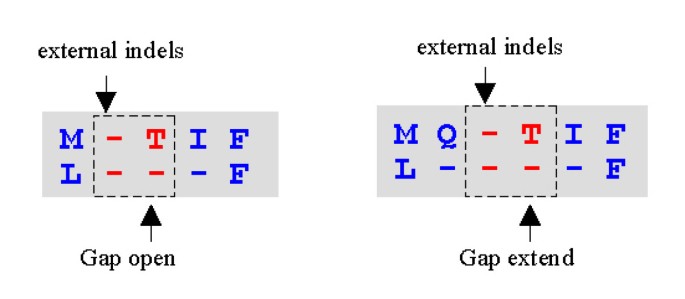
MUSCLE: a multiple sequence alignment method with reduced time and space complexity | BMC Bioinformatics | Full Text